DNA origami nanobots
The designed nanobot is a multifunctional construct with the ability to carry payloads (eg. siRNA, endosomal escape agent), target specific cancer cells, and respond to external stimuli (eg. radio frequency waves). The beauty of nanobot is that it can be modified to carry many different genes in different cancer types by just changing the type of sensor used.
DNA origami technique, where a long scaffold strand is folded into a desired shaped using short ‘staple’ strands, is used to build DNA nano-barrels (NBs) which function as payload carriers. DNA origami technique is chosen as it allows for precise control over the size, shape and functionality of the NBs. The NBs can be designed to express desirable functions, resulting in high efficacy.
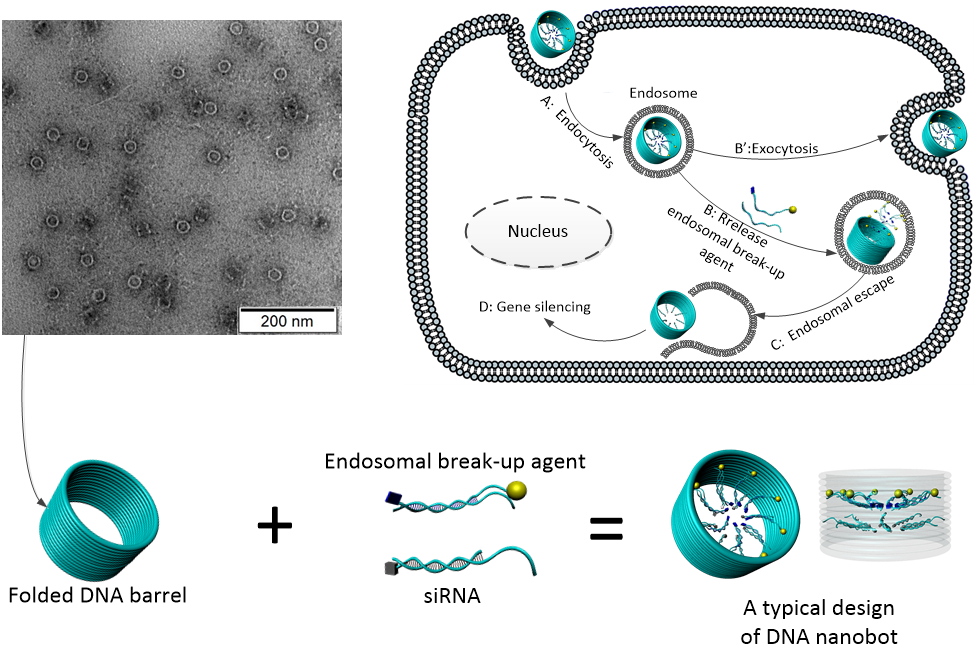
Top left: Nanoscale barrels are self-assembled from DNA using a technique called ‘DNA origami’. Barrels are imaged with a Transmission Electron Microscope (TEM) at the Australian Centre for Microscopy and Microanalysis (ACMM). These barrels are 30 nm in diameter and 30 nm in height.
Top right: Schematic for proposed interaction between a DNA nanobot and a cell. In this project, DNA origami barrels are used to construct a DNA nanobot with RNA aptamer targeting molecules and siRNA cargo. DNA nanobots will be taken up into the endosome, and RF activation will allow endosomal escape.
The interior of these NBs can be functionalized with siRNAs via DNA base pairing. Endosomal escape agents (EEA) are also loaded into the NBs using functionalization using gold nanoparticles as connectors. For targeted delivery of the NBs, the exterior of the NBs are functionalized with cell specific aptamers. Aptamers are single stranded DNA or RNA oligonucleotides capable of binding to a specific target with high affinity. They function like antibodies but are much cheaper and quicker to produce, hence their selection as sensors in this research. These aptamers recognize cancer cell surface markers and allow for receptor-mediated entry of the nanobot into cancer cells.
Upon attachment of the aptamers onto the cancer cells, the entire nanobot is uptaken into the cell via receptor-mediated endocytosis. Upon entry into the endosome, the key challenge is achieving endosomal escape. To achieve this, radio frequency (RF) electromagnetic wave is induced. RF is chosen because it is widely used in thermotherapy and also it can penetrate deeply into the body tissues. Using the right RF, the gold nanoparticle- DNA complex inside the NBs are melted, resulting in the release of endosomal escape agents (EEA) from the NBs. This results in endosomal release of the NBs into the cytoplasm.
The siRNAs are designed in such a way that the RNase III Dicer in the cell dices the siRNA and this detaches the siRNA from the NBs. Upon release from the NBs, siRNA is taken into the RNA-induced silencing complex (RISC) present in the cell, where the duplex siRNA is unwound and the passenger siRNA strand is dissociated. This allows for the siRNA antisense strand to bind to the target mRNA, resulting in subsequent cleavage of the mRNA and silencing of the targeted cancer gene.
Currently, the focus is on the construction of the nanobots. The NBs have been successfully constructed and the next step is the functionalization of the NBs with siRNAs, EEAs and aptamers. Our research group is also currently working on the controlled melting of gold nanoparticle–DNA complex using RF as this is going to be a vital step for endosomal escape. Upon successful construction of the nanobot, in vitro testing will be conducted using specific cancer cell lines.
This case study is a good example of a multidisciplinary research, involving expertise from chemistry, chemical engineering, physics, medicine and biology. The research is a collaboration between the School of Chemical and Biomolecular Engineering, School of Chemistry, and Faculty of Pharmacy.
